Unit 4: Quantification#
Topics: Image analysis, software versions, code, and example availability
Motivation#
Quantitative results are better than qualitative results only. However, it is critical that the performed analysis, as well as the used tools and their settings, are properly documented in the methods. For a custom new analysis, the sequence or code of the performed analysis should be shared.
Key considerations#
Quantifications are better than qualitative results.
Cite software tool and specify version.
Describe workflow.
Disclose critical parameters.
Introduction#
For our example figure, we could easily imagine that we create a quantitative analysis to count the number of living cells vs dead cells. This could either be based on existing tools or writing our own analysis.
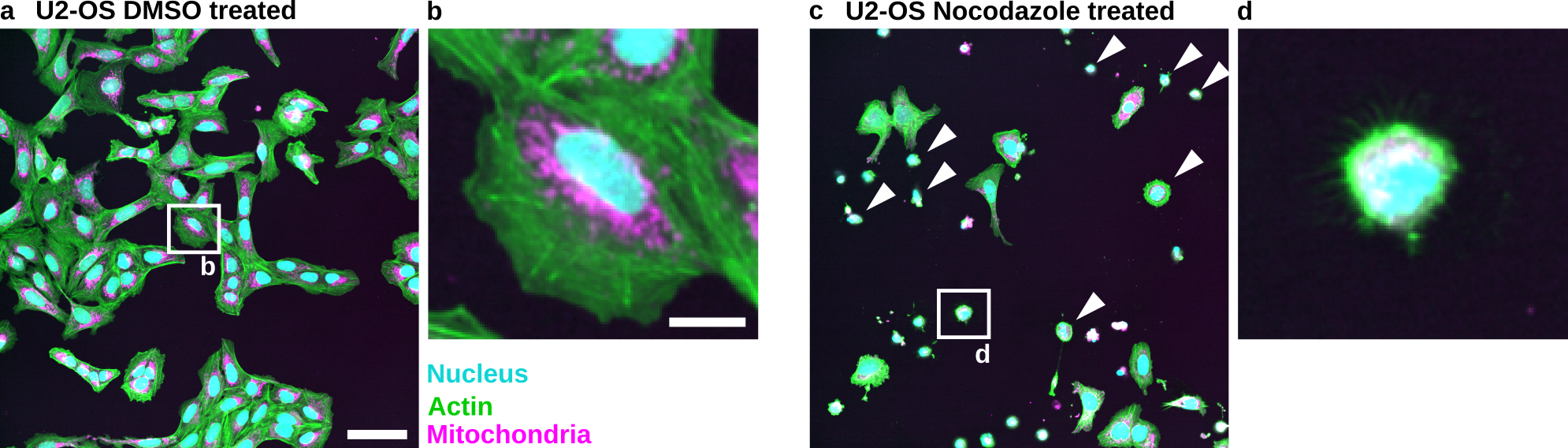
Fig. 54 Nocodazole induces cell death in U2-OS cells: (a) U2-OS cells treated with DMSO only. (b) Cropped image of a single cell treated with DMSO. (c) U2-OS cells treated with Nocodazole at a 5 µM concentration show increased cell death. Note the many small round cells (White Arrowheads). (d) Cropped image of a dead or dying cell. Scale bar represents 100 µm (a) and 20 µm (b).#
Download the example images here:
Analysis based on existing workflows#
For our example images, I was able to adopt one of my own published tools, OrgaMapper:
Schmied, C., Ebner, M., Samsó, P. et al. OrgaMapper: a robust and easy-to-use workflow for analyzing organelle positioning. BMC Biol 22, 220 (2024). https://doi.org/10.1186/s12915-024-02015-8
To reproduce the analysis, you will need to get the plugin in Fiji:
Help > Update…
Fig. 55 Update Fiji.#
Press: Manage update site
Select Cellular Imaging Facility
Press ok and restart Fiji
Save the example images in an input folder (e.g., test). OrgaMapper documents the analysis settings using an .xml file. Please download the settings: settings .xml and put it into the input folder. Create an output folder (e.g., test_out).
Start OrgaMapper:
Plugins > CellularImaging > Map Organelle
Specify the location of the input folder, output folder, and the settings file. Then press ok to start OrgaMapper.
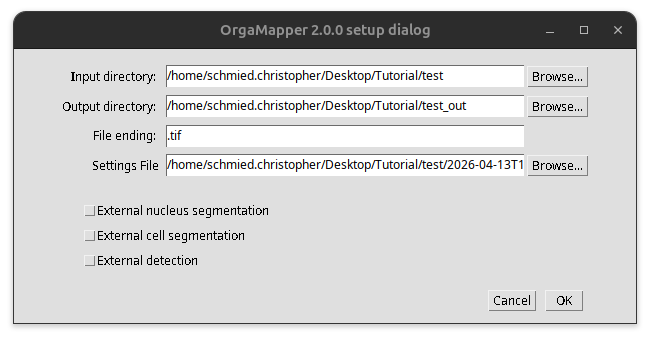
Fig. 56 Start OrgaMapper.#
OrgaMapper then allows you to perform a simple watershed-based image segmentation and perform a basic analysis of each cell over the control and treatment. For more information about the analysis performed, you can go to the documentation of OrgaMapper.
OrgaMapper visualizes this on the example images using the “Preview” button:
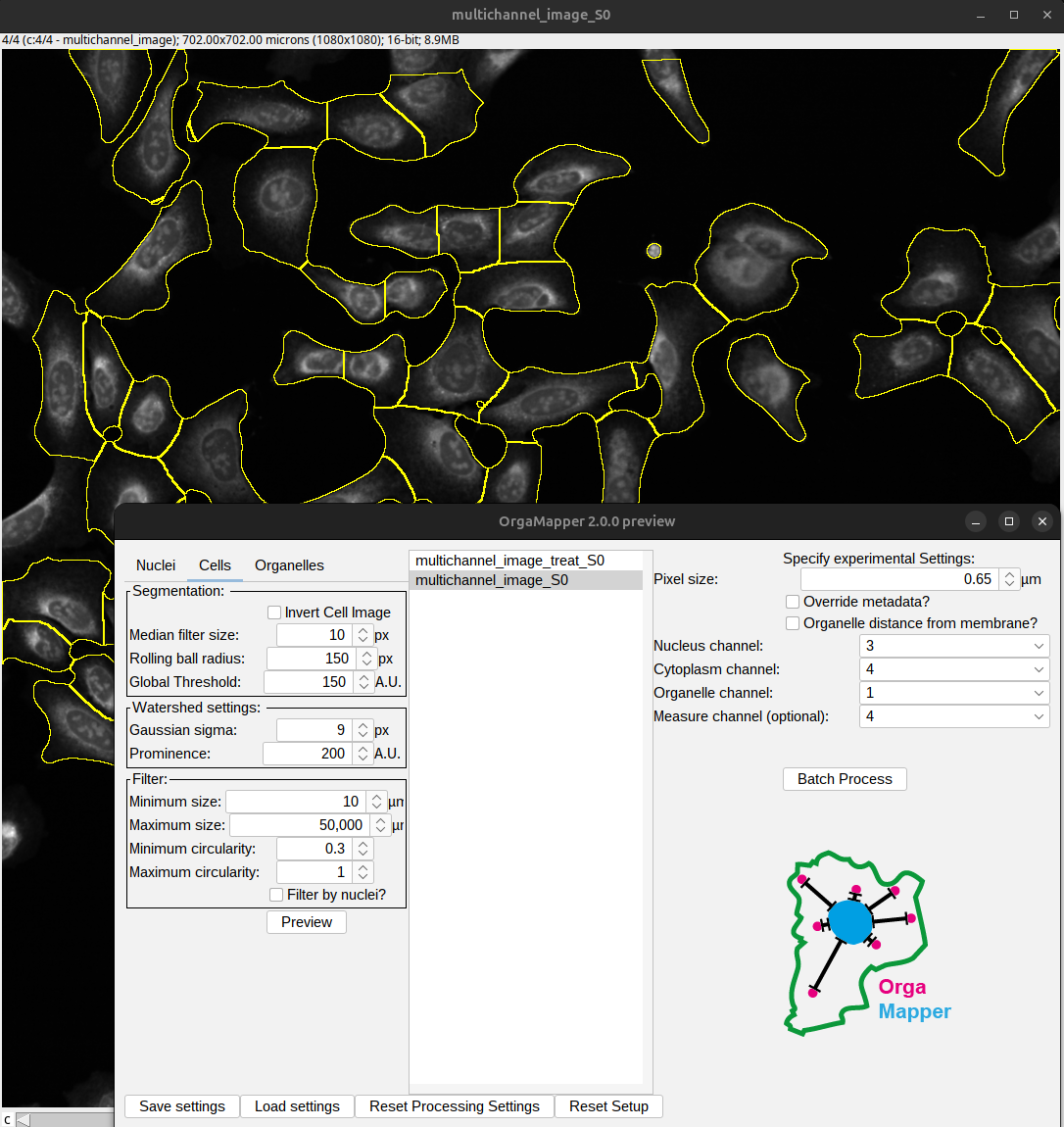
Fig. 57 Start OrgaMapper.#
To execute the analysis, press the “Batch Process” button. OrgaMapper then performs the analysis based on the loaded settings and will save the results in the output folder.
Fig. 58 OrgaMapper Results.#
Critical is that we can extract the cell area in the “cellMeasurements.csv” file:
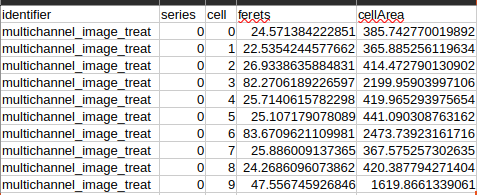
Fig. 59 OrgaMapper cell area results.#
I then performed a simple threshold, asking how many cells in the control image, as well as in the Nocodazole treated image, are above or below 1000 µm². As a proxy for assessing cell death. Which yields the following result:
Treatment |
Cells with area <1000 µm² |
Cells with area >1000 µm² |
Total cells |
% dead cells |
|---|---|---|---|---|
DMSO |
0 |
63 |
63 |
0 |
Nocodazole |
23 |
6 |
29 |
79 |
Note
This example was quickly created to illustrate the point. It does not claim to be correct or a good analysis of cell death.
This quantification could now be in the result of your manuscript. It is important to properly document the performed analysis. For this, it is again critical to document the used software platform as well as the tools and their versions:
Image analysis was performed using Fiji is just ImageJ (Fiji) (Schindelin et al. 2012) version: 2.16/1.54p. and the OrgaMapper plugin version 2.0.0 (Schmied et al. 2024). Nuclei were segemented using an intensity based segmentation using the Li automatic threshold after filtering with a median filter with size 5 pixels and a rolling ball background subtraction with radius of 10 pixels. After thresholding, an erosion of 1 was applied to the Nuclei masks, and the segmented nuclei were filtered based on size, with nuclei kept between 1 µm² and 750 µm². The cell area was segemented using an intensity based threshold with a manual threshold of 150 Fluorescent Intensity (A.U.) after a median filter was applied with a filter size of 10 pixels and a rolling ball background subtraction with a radius of 150 pixels. For the watershed-based cell segmentation, the centers of cells were detected using a summed nuclei and cytoplasm channel. To detect the cell centers, a Laplacian-of-Gaussian filter was applied with a sigma of 2 and peak values with a prominence of 40 Fluorescent Intensity (A.U.). Separated cells were then filtered by keeping cells above 10 µm² and a circularity above 0.3. For each cell, the cell area was further analyzed. Dead cells were defined with a threshold below 1000 µm².
Schindelin, J., Arganda-Carreras, I., Frise, E. et al. Fiji: an open-source platform for biological-image analysis. Nat Methods 9, 676–682 (2012). https://doi.org/10.1038/nmeth.2019
Schmied, C., Ebner, M., Samsó, P. et al. OrgaMapper: a robust and easy-to-use workflow for analyzing organelle positioning. BMC Biol 22, 220 (2024). https://doi.org/10.1186/s12915-024-02015-8
Tip
Since OrgaMapper provides a settings file, this settings file can be shared in a data repository.
Analysis based on new workflows#
For this analysis, we could easily also write a custom code based on a Fiji macro. To properly document such an analysis, we should include the code and example data in a data repository for review.
Code and example availability#
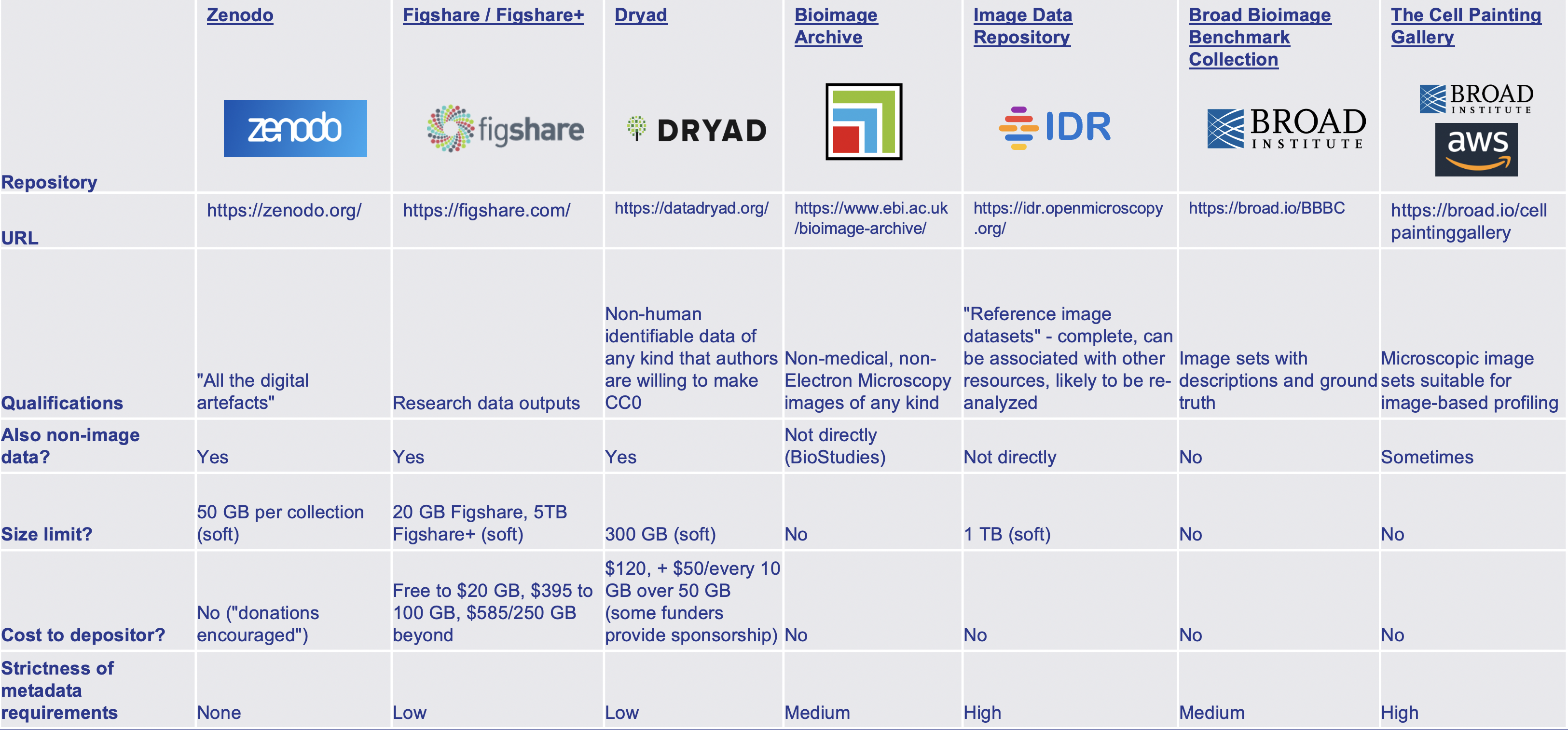
Fig. 60 Examples of different repositories. Overview provided by Cimini 2023.#